Projects
Morloc
morloc is a functional programming language where
functions are imported from foreign languages and unified through a common type
system. The compiler generates the code needed to compose functions across
languages and also to direct automation of mundane tasks such as data
validation, type/format conversions, data caching, distributed computing, and
file reading/writing. The ultimate goal is to develop morloc into a query
language that returns optimized programs from an infinite “library” of
functions and compositions of functions.
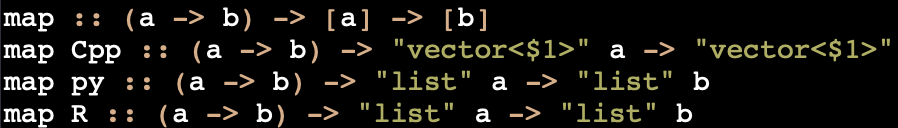
Import many implementations for a function from any supported language:

Declare one common type signature that describes many implementation-specific ones
Specify new programs by composing these general functions

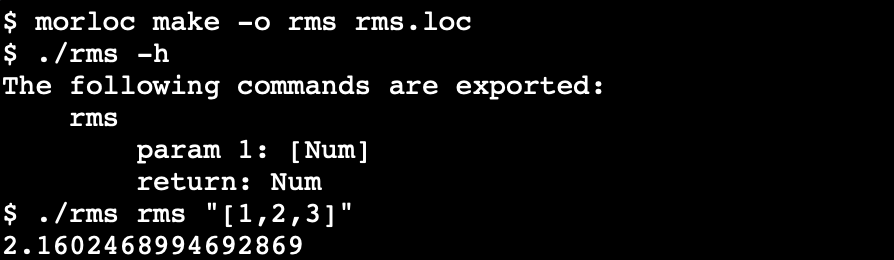
Build executables and let the compiler optimize the choice of implementations
Haskell projects
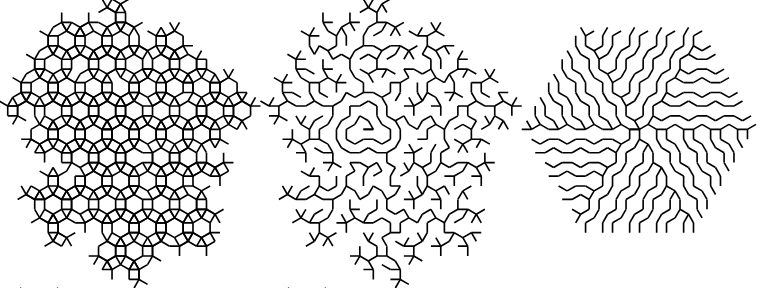
- lsystems an experimental package for specifying and generating deterministic or stochastic L-systems graphics.

- tessellate an experimental semi-regular tesselation package with aspirations of growing up to be something different.

Python packages on PyPi
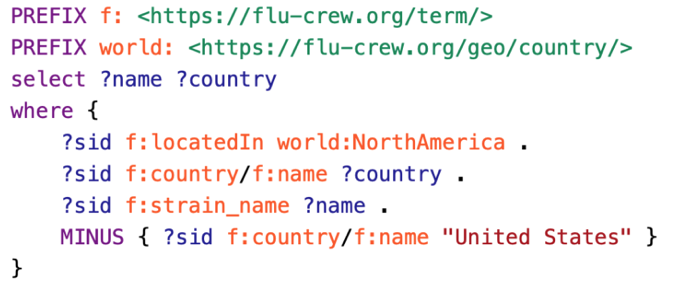
pgraphdb is a Python library and CLI tool for interacting with a GraphDB database. It allows uploading of RDF data, submission of SPARQL queries, deletion of triples specified in RDF files, etc.
octofludb is a specialized tool for parsing the data used in our swine influenza surveillance program into triple format and uploading it to a GraphDB database. It allows access to the data through SPARQL queries and can return FASTA file sequence.
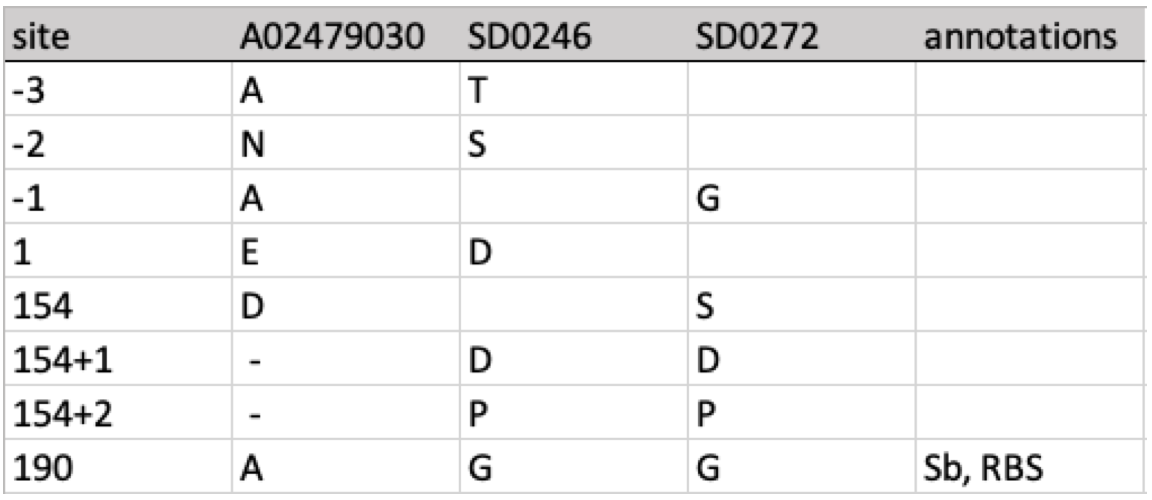
- flutile is a specialized suite of tools designed for the flu-crew at USDA-ARS and collaborators. The foundational feature is handling for indexing amino acid positions relative to references for each influenza A subtype. This allows sequences to be compared across studies.
- smot is a python package and CLI tool for general purpose phylogenetic tree sub-sampling, annotation, and summarization.

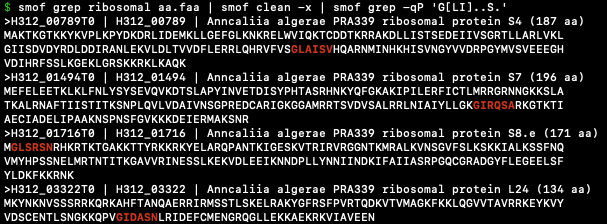
- smof is a CLI tool for working with FASTA files.
- regref offers regular expression based search and replace using pattern and replacement expressions from a table.
R CRAN/rOpenSci packages
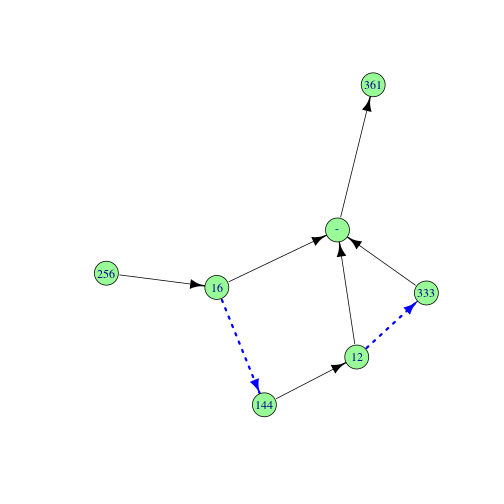
- rmonad an experimental monadic pipeline tool for building branching workflows, storing select intermediate data, automatic benchmarking, and storing of any raised error messages or warnings.

rhmmer a wrapper around the bioinformatics package HMMER.
onekp scrape plant transcritomic data from the 1KP project website, select the species or clades or interest, and automatically download them.
Other
- Language Of Dice was my first attempt at developing a programming language. The goal was to develop a language for specifying discrete probability problems (dice problems) that was sufficiently advanced to model outcomes of encounters in D&D. For example, what is the probability that my group of 3 level-2 fighters will beat the Kraken. All possible attacks, buffs, debuffs, etc would have to be modeled. The project currently consists only of loose specification of syntax, grammar, and a rudimentary lexer and parser.
R Shiny apps
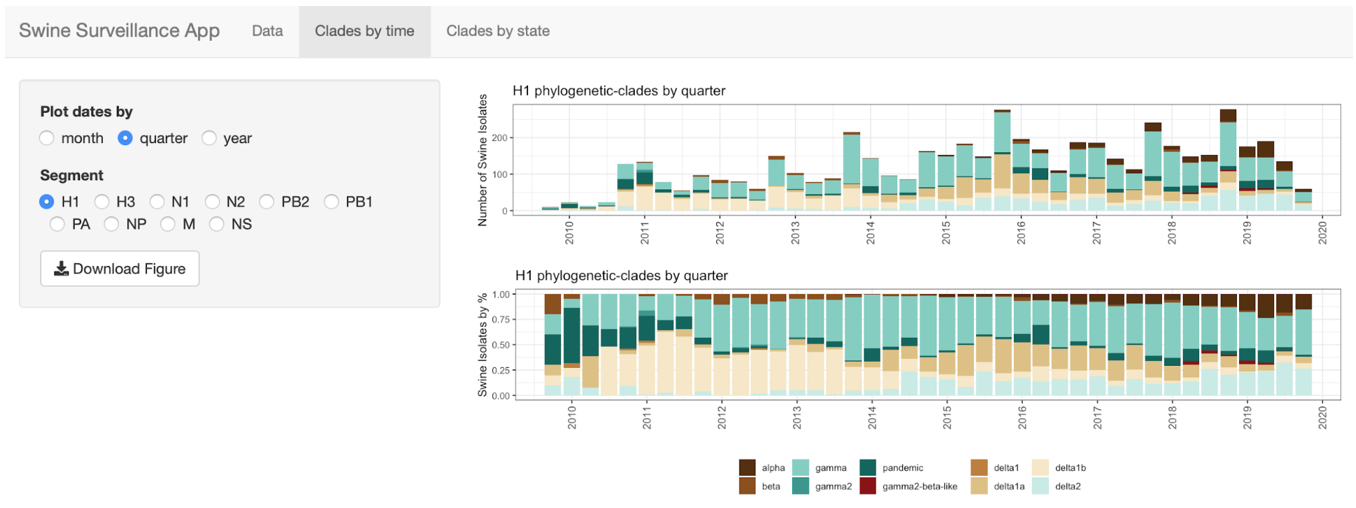
octoflushow (private) imports data
generated by octofludb and produces interactive visuals and tables for
subsetting/downloading data.
metaoku (app) was my grand vision for a means of sharing and visualizing data organized into normal folders. The project is described in my dissertation appendix. The project is basically dead now, but it seemed my approach to automatically visualizing data and my simlple data type-system are worth returning to at some point.
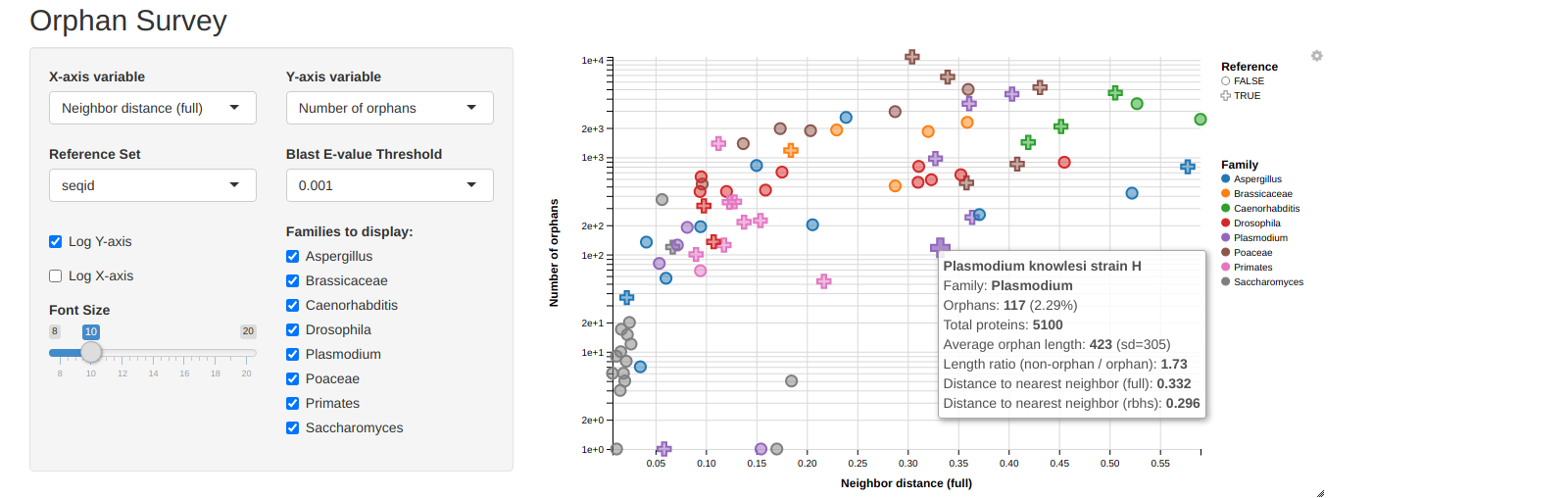
Orphan survey (app) is a Shiny visualization program for an unpublished orphan gene study I ran in grad school. Nothing come of it ultimately.
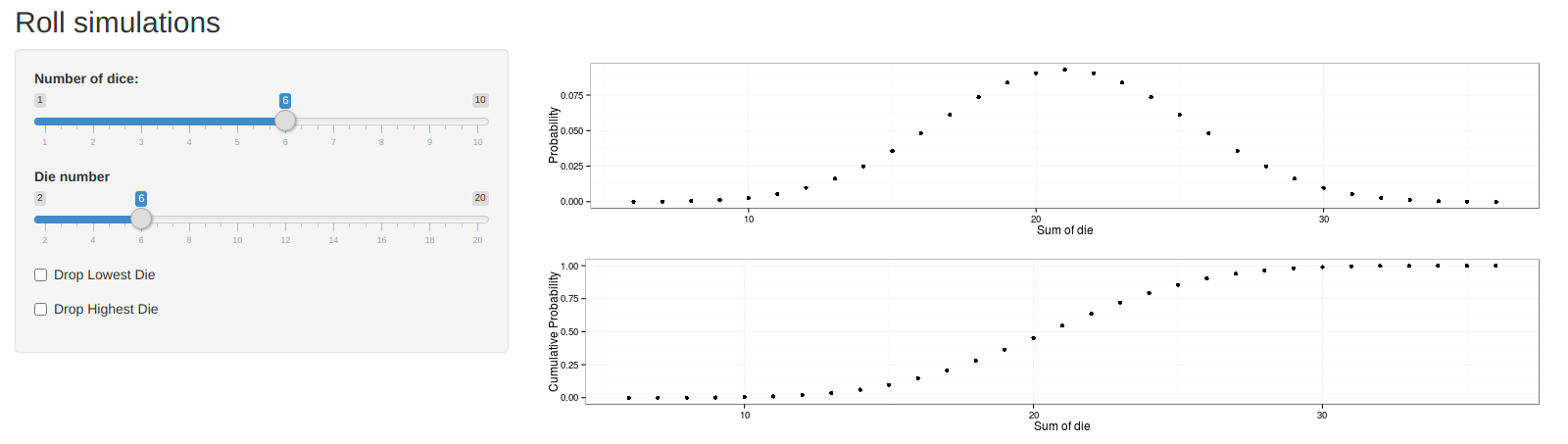
dnd (app) is a rudimentary dice probability app. Someday, when I feel sufficiently motivated, I’ll come back to it and expand the functionallity. Dice probability gets interesting when you start adding in advantage/disadvantage and other D&D shenanigans.
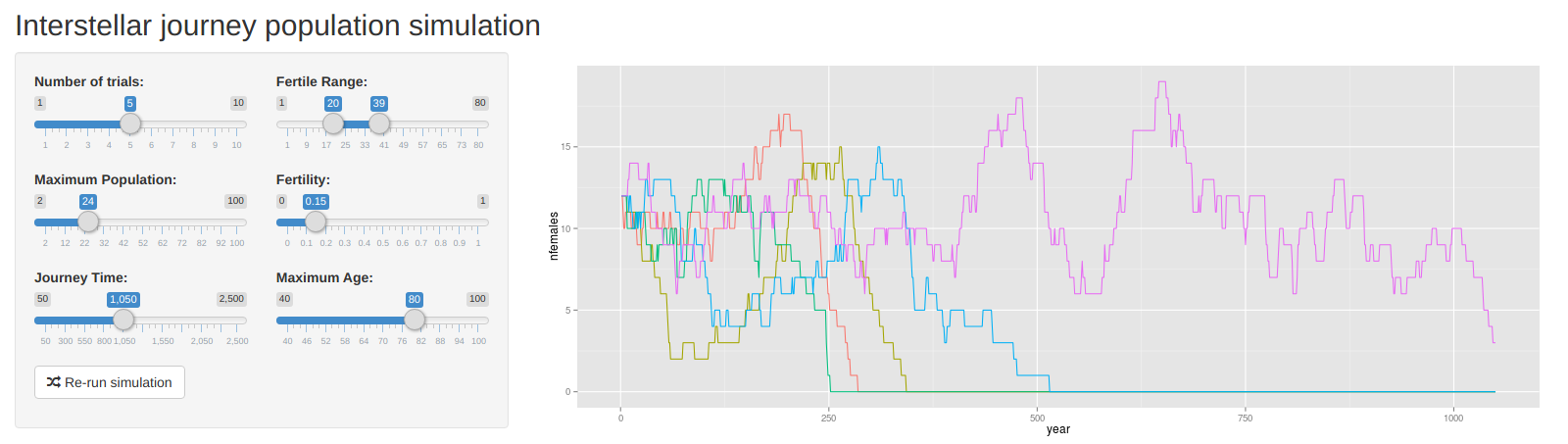
Small population survival (app) was a visualization app for a class project concerning how long a colony on a generation ship would survive before the proportion of females reched 0 or 1 and natural reproduction became impossible.
Genomics and orphan-gene sleuthing from grad-school
- synder is a high-performance program that maps genomic loci from one species to another using a synteny map. The core algorithm is written in C++ and wrapped in R.

- phylostratr an R pipeline for conventional phylostratigraphy.

- fagin an R pipline based on
synderandrmonadfor inferring the origins of genes.
